The Zhang Translational Genomics Laboratory develops and applies multi-omics and deep learning approaches in cancer biology and personalized radiation oncology. We collaborate with researchers and physician scientists from Department of Radiation Oncology, McDonnell Genome Institute, Institute for Informatics, Siteman Cancer Center, and other Washington University research labs and national and international institutes.
My previous research interests included computational biology and genomics algorithms as they apply to translational cancer research. My doctoral work focused on developing structural variation discovery tools using next-generation sequencing data, including SVseq 1 & 2. I also worked on algorithmic problems in haplotype inference, recombination, rare variants, etc. My postdoctoral work focused on developing algorithms analyzing whole transcriptome sequencing data to discover RNA specific aberrations and their applications in cancers. We designed and implemented the state-of-the-art gene fusion discovery tool, INTEGRATE, leading to the discoveries of novel biomarkers in breast cancer, liver cancer, leukemia, etc. We implemented the first tool in cancer immunology, INTEGRATE-Neo, to predict neo-antigens from tumor specific gene fusion peptides. We discovered a single-gene biomarker, lncRNA PCAT-14, in prostate cancer metastasis, and novel mid-sized small RNAs in acute myeloid leukemia and prostate cancer.
We are currently working on creating and integrating deep learning and radiogenomics approaches using multi-omics data (RNA-seq, scRNA-seq, WGS, proteomics, metabolomics) and imaging (PET/CT/MRI) data into the development of novel diagnostic, prognostic, and therapeutic strategies in cancers. My long-term goal is to leverage my unique training and expertise in computer science, data science, cancer genomics, population health sciences, and radiation oncology to create novel computational approaches using muti-omics and longitudinal data (i.e., genomics, proteomics, metabolomics, imaging, and clinical) to interpret how molecular alterations in cancers affect patient responses to therapies. These translational deep learning models, and radio-genomic and multi-omics analyses, will ultimately facilitate the prevention, diagnosis, and treatment of cancers and improve patient outcomes.
Ongoing projects:
NCI R01 Research Project Grant – Title: Integrating multi-omics, imaging, and longitudinal data to predict radiation response in cervical cancer. $1,663,760
NCI R37 Research Project Grant (Converted from R01) – Title: HPV genomic structure in cervical cancer radiation response and recurrence detection. $1,621,283
NCI ITCR R21 Developmental Research Grant Award – Title: Deep learning in cervical cancer radiogenomics. $257,125
NCI K22 Transition Career Development Award – Title: HPV alternative splicing in cervical cancer radiation response. $438,661
Recently finished projects:
ICTS Clinical and Translational Research Funding Program Award – Title: Examining cervical cancer HPV genotypic radiation response using augmented structural gene expression differences. $50,000
SIP Pre-R01 multi-PI Award – Title: HPV genomic structural subtypes in oropharyngeal squamous cell carcinoma. $200,000
Research Areas:
I. Machine learning/ Deep learning/ and Radiogenomics
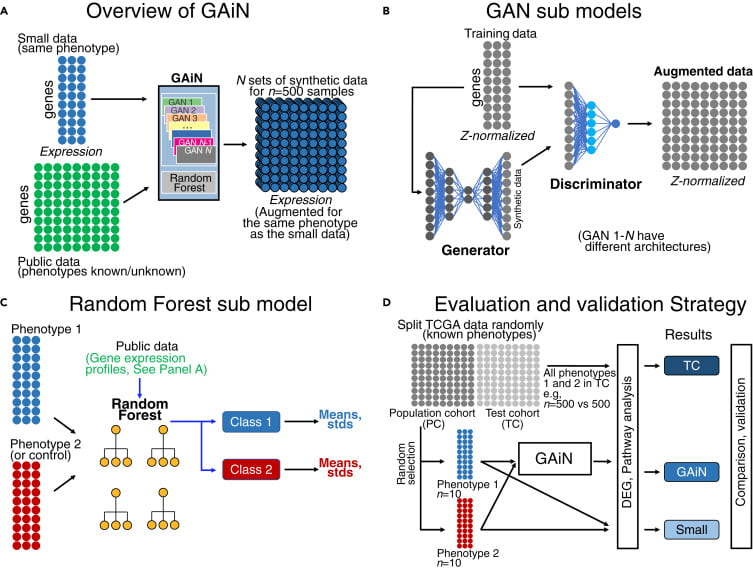
ML and DL is one of our lab’s most current research focuses. Please refer to News for ongoing grants and projects.
https://www.linkedin.com/feed/update/urn:li:activity:7150901702462689280/

Tumor Evolution (Science Advances 2020); SVM in SV (PLoS One 2014); Synchronization Detection (BMC Genomics 2013); Probabilistic method (BMC Genomics 2013); Logic Regression (ISCABS 2011); Haplotype Inference (PSB 2011)
II. Algorithms and tools
INTEGRATE: gene fusion discovery using whole genome and whole transcriptome data

While next-generation sequencing (NGS) has become the primary technology for discovering gene fusions, we are still faced with the challenge of ensuring that causative mutations are not missed while minimizing false positives. Currently, there are many computational tools that predict structural variations (SV) and gene fusions using whole genome (WGS) and transcriptome sequencing (RNA-seq) data separately. However, as both WGS and RNA-seq have their limitations when used independently, we hypothesize that the orthogonal validation from integrating both data could generate a sensitive and specific approach for detecting high-confidence gene fusion predictions. Fortunately, decreasing NGS costs have resulted in a growing quantity of patients with both data available. Therefore, we developed a gene fusion discovery tool, INTEGRATE, that leverages both RNA-seq and WGS data to reconstruct gene fusion junctions and genomic breakpoints by split-read mapping. To evaluate INTEGRATE, we compared it with eight additional gene fusion discovery tools using the well-characterized breast cell line HCC1395 and peripheral blood lymphocytes derived from the same patient (HCC1395BL). The predictions subsequently underwent a targeted validation leading to the discovery of 131 novel fusions in addition to the seven previously reported fusions. Overall, INTEGRATE only missed six out of the 138 validated fusions and had the highest accuracy of the nine tools evaluated. Additionally, we applied INTEGRATE to 62 breast cancer patients from The Cancer Genome Atlas (TCGA) and found multiple recurrent gene fusions including a subset involving estrogen receptor. Taken together, INTEGRATE is a highly sensitive and accurate tool that is freely available for academic use.
Zhang J, et al. Genome Research 2016. PMID: 26556708
HPV-EM: an accurate HPV detection and genotyping EM algorithm

Accurate HPV genotyping is crucial in facilitating epidemiology studies, vaccine trials, and HPV-related cancer research. Contemporary HPV genotyping assays only detect < 25% of all known HPV genotypes and are not accurate for low-risk or mixed HPV genotypes. Current genomic HPV genotyping algorithms use a simple read-alignment and filtering strategy that has difficulty handling repeats and homology sequences. Therefore, we have developed an optimized expectation–maximization algorithm, designated HPV-EM, to address the ambiguities caused by repetitive sequencing reads. HPV-EM achieved 97–100% accuracy when benchmarked using cell line data and TCGA cervical cancer data. We also validated HPV-EM using DNA tiling data on an institutional cervical cancer cohort (96.5% accuracy). Using HPV-EM, we demonstrated HPV genotypic differences in recurrence and patient outcomes in cervical and head and neck cancers.
Inkman M. et al. Scientific Reports 2020. PMID: 32868873
More algorithms and tools:
INTEGRATE (Genome Research 2016); INTEGRATE-Neo (Bioinformatics 2017); INTEGRATE-Vis (Scientific Reports 2017); SVseq2 (BMC Bioinformatics 2012); SVseq(Bioinformatics)
III. Applications of gene fusions and structural variations
Functional Annotation of ESR1 Gene Fusions in Estrogen Receptor-Positive Breast Cancer


ESR1 gene fusions promote endocrine therapy resistant cell proliferation and metastasis.
More on gene fusions and SVs
SMC-RNA Dream challenge (Cell Systems 2021) ESR1 EMT(Cell Reports 2018); ESR1 Treatment Resistance (Cell Reports 2013); Prostate Cancer (Cell 2018); mFL-HCC (Annals of Oncology 2016); Adult B-lymphoblastic leukemia (Experimental Hematology 2016)
IV. HPV-related cancer biology, metabolism, and radiogenomics
Integrating imaging and RNA-seq improves outcome prediction in cervical cancer

Approaches using a single type of data have been applied to classify human tumors. Here we integrate imaging features and transcriptomic data using a prospectively collected tumor bank. We demonstrate that increased maximum standardized uptake value on pretreatment 18F-fluorodeoxyglucose-positron emission tomography correlates with epithelial-to-mesenchymal transition (EMT) gene expression. We derived and validated 3 major molecular groups, namely squamous epithelial, squamous mesenchymal, and adenocarcinoma, using prospectively collected institutional (n = 67) and publicly available (n = 304) data sets. Patients with tumors of the squamous mesenchymal subtype showed inferior survival outcomes compared with the other 2 molecular groups. High mesenchymal gene expression in cervical cancer cells positively correlated with the capacity to form spheroids and with resistance to radiation. CaSki organoids were radiation-resistant but sensitive to the glycolysis inhibitor, 2-DG. These experiments provide a strategy for response prediction by integrating large data sets, and highlight the potential for metabolic therapy to influence EMT phenotypes in cervical cancer.
Zhang J, et al. Journal of Clinical Investigation 2021. PMID: 33645544
From QuadShot News: A hot mes | Zhang, J Clin Invest 2021
More on cervical cancer biology, metabolism, and radiogenomics:
HPV and CRT (JCI Insight 2021); SUVmax and Macrophage (Clinical Cancer Research 2021); Glutaminase Inhibitors (Molecular Cancer Therapeutics 2020); Neutrophils (PNAS 2019); SUVmax/Radiogenomics (ASTRO 2018)
V. Non-coding RNAs
Comprehensive discovery of noncoding RNAs in acute myeloid leukemia cell transcriptomes

Zhang J, et al. Experimental Hematology 2017. PMID: 28760689
Featured in: RNA Biology Blog
Multi-institutional Analysis Shows that Low PCAT-14 Expression Associates with Poor Outcomes in Prostate Cancer

Integrative analysis reveals Prostate Cancer Associated Transcript-14 (PCAT-14) expression associates with prostate cancer
More on non-coding RNA:
PCAT-14 (European Urology 2017); Mid-sized RNA (Experimental Hematology 2017); cDNA Capture (The Journal of Molecular Diagnostics 2014)
VI. Population genetics and cancer evolution
Tumor Evolution (Science Advances 2020); Rare Variants (BMC Genomics 2013); Haplotype Inference (PSB 2011); SNP and Logic Regression (ISCABS 2011)
VII. Cancers, immunology, radiation therapy, and other clinical applications
Cervical cancer (JCI Insight 2021; Clinical Cancer Research 2021; Journal of Clinical Investigation 2021; Molecular Cancer Therapeutics 2020; Scientific Reports 2020 ; PNAS 2019; ASTRO 2018) Breast cancer (Cell Reports 2018; Genome Research 2016; Cell Reports 2013) Prostate Cancer (Journal for ImmunoTherapy of Cancer 2020; Cell 2018; Bioinformatics 2017; Scientific Reports 2017; European Urology 2017; The Journal of Molecular Diagnostics 2014) Colorectal cancer (Science Advances 2020) Head and neck cancer (Scientific Reports 2020) Leukemia (Experimental Hematology 2017; Experimental Hematology 2016) Lung cancer (The Journal of Molecular Diagnostics 2014) mFL-HCC (Annals of Oncology 2016) Immunology (Journal for ImmunoTherapy of Cancer 2020; Bioinformatics 2017)
Please also refer to the full list of publications.