
Welcome to the Djuranovic Lab!
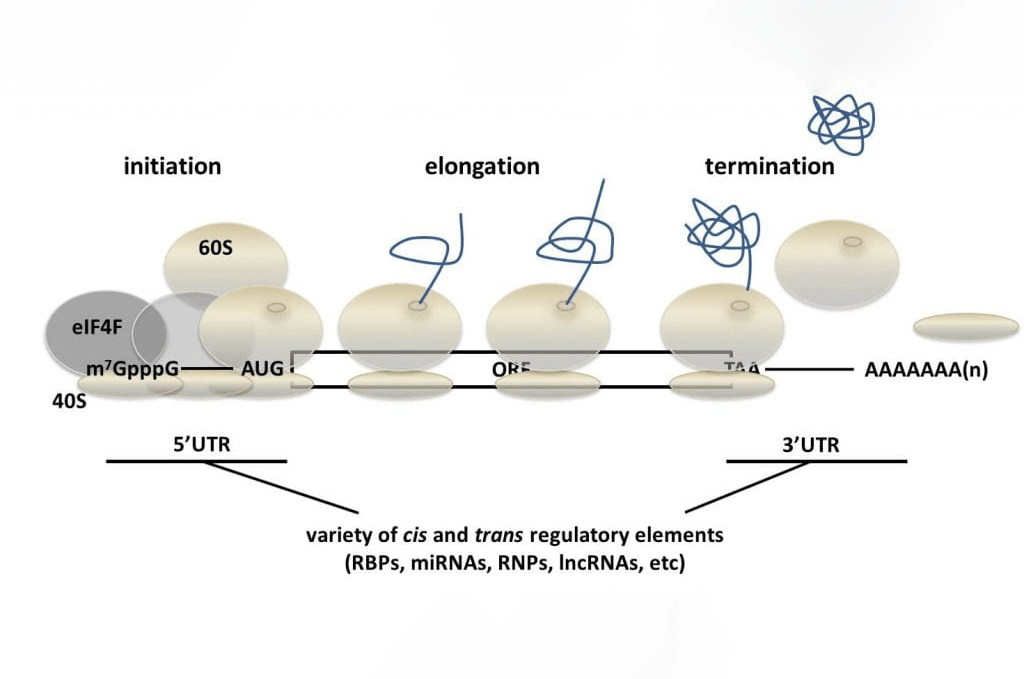
The Djuranovic Lab is primarily interested in understanding the mechanisms of post-transcriptional gene regulation. We focus on RNA-binding proteins (RBPs), ribonucleoprotein (RNP complexes) as well as mRNA sequence motifs, all of which control translational efficiency of their target mRNAs or are involved in general RNA metabolism.
Explore our research!

Meet the team!
Latest News from Djuranovic Lab
Two former Djuranovic lab members started (October 1st 2022) or will start (January 1st of 2023) their independent labs. Good luck and lot of success in your future work!!!
Slavica Pavlovic Djuranovic (Assistant Professor at Cell Biology Department at WashU)
https://cellbiology.wustl.edu/people/pavlovic-djuranovic/
Kyle Cottrell ( Assistant Professor at Biochemistry Department at Purdue University)
https://www.cottrellrna.com/
We are happy to announce that Sergej was promoted to full professor of Cell Biology and Physiology at Washington University School of Medicine starting January 1st 2023.

Another collaborative effort got funded for Djuranovic Lab as well as Dougherty, Miller and Kroll Labs at WashU School of Medicine. We will be looking for treatment of MYT1L disruption that leads to ASD…more news on this soon
The CZI funds for treatment of neurodegeneration are here. We are looking for a graduate students and postdoctoral fellows to work on this and other projects.

@DjuranovicLab and @slavicaPD lab traditionally presenting at Cold Spring Harbor Translation Control meeting




Collaborative study from Zaher lab and @DjuranovicLab is out in Cell Reports, check it out: N1-methylpseudouridine found within COVID-19 mRNA vaccines produces faithful protein products
https://www.sciencedirect.com/science/article/pii/S2211124722011202

Great news for @MillerLabNeuro and @DjuranovicLab from @czi. Our project on treating neurodegeneration has received funds for the next 4 years. A big thanks to Chan Zuckerberg Initiative and Neurodegeneration Challenge Network on their support.
https://chanzuckerberg.com/science/programs-resources/neurodegeneration-challenge/projects/
Collaborative study from Zaher lab and @DjuranovicLab is out in Cell Reports, check it out: N1-methylpseudouridine found within COVID-19 mRNA vaccines produces faithful protein products:
https://t.co/Dx3JtQ44fi

Djuranovic Lab was presenting novel findings at the 2022 Ribosome meeting in Bordeaux


Science is also fun – new @DjuranovicLab members with a little help of @slavicaPD are being creative using E. coli, couple of different plasmids and LB plates with arabinose. More decorations with varying amounts of fluorescent proteins or frameshifted reporters soon…

New manuscript from Djuranovic lab is out in Journal of Biological Chemistry. The manuscript is a part of Jessey’s PhD thesis. Jessey shows how RACK1 protein interacts with ribosomes in human and P. falciparum:
https://www.sciencedirect.com/science/article/pii/S0021925822003945

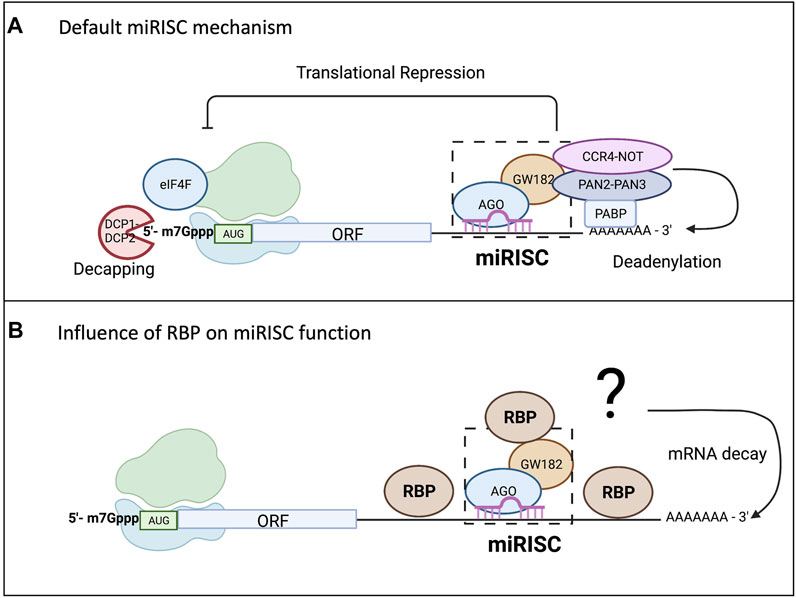
Check our new article on the role of RNA-binding proteins in modulation of microRNA-mediated gene regulation. Courtney gives a nice overview of what is known and what still needs to be answered on this topic:
https://www.frontiersin.org/articles/10.3389/fmolb.2022.832916/full
Geralle’s new paper on use of polyA tracks in combination with CRISPR/Cas9 technology is out in Molecular Therapy. You can check it on this link:
https://www.cell.com/molecular-therapy-family/nucleic-acids/fulltext/S2162-2531(21)00248-1

We are grateful to @NIH and @NIGMS for providing their support on our studies of the mRNA surveillance and mRNA-ribosome interactions in AU-rich genome of P. falciparum. We just got official NOA for our R01. Thanks to collaborators in Goldberg, Puglisi and Jovanovic labs.

We are happy to host three rotation students and 2 undergraduate researchers in our lab this year.
Rotation students from MCB program are Jade Enright, Katherine Floyd and Sarah Koester, undergraduate researchers are Eylül Horozoğlu and Anika Gupta.
New manuscript on usage of polyA track method for gene dosage control and short study on AUF1 and TP53 hypomorphs from our lab. Gene dosage effects of polyA track engineered hypomorphs by Geralle Powell.

https://www.biorxiv.org/content/10.1101/2021.02.10.430645v1
We are thankful to @SitemanCenter for sponsoring our research on polyA tracks, mutations and cancer.

Dr. Jessey Erath is the new PhD from Djuranovic Lab. Jessey presented his thesis work on Plasmodium falciparum translation machinery and how malaria causing parasite deals with long polyA tracks.

Djuranovic Lab is thankful NIGMS and NIH for continuing support for our work on “Modulation of miRNA mediated translational repression by RBPs and RNA. We have successfully renewed R01 for this project. Expect more results in this direction.

Djuranovic and Miller lab got Chan Zuckerberg Initiative 2020 Collaborative Pilot Project Award for the use of Antisense Oligonucleotides in the treatment of ALS and FTD. More information on the link below.
Geralle Powell, PhD candidate from Djuranovic Lab, will talk about her work on human polyA track genes and Nonsense Mediated Decay (NMD) at this year Cold Spring Harbor Meeting on Translational control. The title of her talk will be:
“NMD regulates expression of endogenous polyA track genes”

New manuscript from Djuranovic lab is out in the eLife. See first part of the story on how malaria causing parasites synthesize poly-lysine from polyA repeats.
https://elifesciences.org/articles/57799

Manasvi receives 2020 Spector Prize for her honor thesis. Each year, the Department of Biology in Arts & Sciences at Washington University in St. Louis awards a prize to a graduating senior in memory of Marion Smith Spector, a 1938 graduate who studied zoology under the late Viktor Hamburger. The Spector Prize, first awarded in 1974, recognizes academic excellence and outstanding undergraduate achievement in research.
https://biology.wustl.edu/news/manasvi-verma-wins-2020-spector-prize

While Manasvi is still choosing her future graduate school. We got this notification from Nature Communications.

We have a new member of the lab – Courtney Jungers.

Courtney is a MCB program PhD candidate and she will be working on topics of translational control by miRNAs and RBPs.
Manasvi’s and Kyle’s paper has been seen more than 12000 times in less than a month of being published and it was picked up by 20 news outlets.
https://www.nature.com/articles/s41467-019-13810-1/metrics

Manasvi’s and Kyle’s paper is out.
Verma, M., J. Choi, K. A. Cottrell, Z. Lavagnino, E. N. Thomas, S. Pavlovic-Djuranovic, P. Szczesny, D. W. Piston, H. S. Zaher, J. D. Puglisi and S. Djuranovic (2019). “A short translational ramp determines the efficiency of protein synthesis.” Nature Communications 10(1): 5774. https://doi.org/10.1038/s41467-019-13810-1
It was highlighted in the news.
https://medicine.wustl.edu/news/scientists-find-way-to-supercharge-protein-production/

Our review on polyA tracks in malaria causing parasite Plasmodium falciparum is out.
Erath, J., S. Djuranovic and S. P. Djuranovic (2019). “Adaptation of Translational Machinery in Malaria Parasites to Accommodate Translation of Poly-Adenosine Stretches Throughout Its Life Cycle.” Frontiers in Microbiology 10: 2823. https://doi.org/10.3389/fmicb.2019.02823
